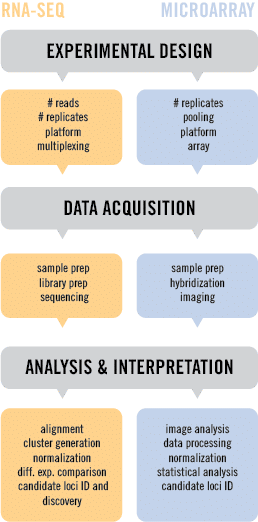
Since it’s inception, RNA-seq has been compared to microarray technology as a means of generating transcriptome information. Both follow a similar path to answering a biological question. This begins with experimental design, followed by data acquisition, and finally analysis and interpretation. However, there are a few key differences between the technologies, and here at Cofactor, we believe that RNA-seq has a number of advantages.
First, using microarray technology limits the researcher to detecting transcripts that correspond to existing genomic sequencing information. RNA-seq experiments on the other hand work well for investigating both known transcripts and exploring new ones. Therefore, RNA-seq is ideal for discovery-based experiments. And while microarrays are limited to the reference information available during production, RNA-seq experiments may be updated as new sequence information is obtained.
Second, RNA-seq delivers low background signal. This is because DNA sequences can be unambiguously mapped to unique regions of the genome. As a result, noise in the experiment is easily eliminated during analysis. Hybridization issues seen with microarrays, such as cross-hybridization or non-ideal hybridization kinetics, are also eliminated in RNA-seq experiments. This offers another signal-to-noise advantage.
Finally, RNA-seq has the ability to quantify a large dynamic range of expression levels, with absolute rather than relative values. Even with organisms lacking a reference genome, de novo transcriptome assembly and differential expression analysis can be performed. There is no upper limit for this quantification, and at Cofactor, we can help you to determine the number of reads you need to find the information you’re looking for with high levels of reproducibility between both technical and biological replicates. What’s more, the statistical analysis we perform can add an additional level of confidence to your results.
Based on current technology, in most cases, the upfront cost of performing a microarray-based experiment will cost less than an RNA-seq based experiment, and may be the most appropriate solution, based on your goals. However, if the objectives of your experiment hinge on the three important considerations highlighted above (sensitivity, discovery, range of expression) RNA-seq is going to be your best bet. This will end up being cheaper and more time efficient than starting with microarrays and having to end up using RNA-seq later anyway. By now, most researchers interested in studying gene expression and the dynamic landscape of the transcriptome have fully adopted RNA-seq for this very reason as well as many others, not including the aforementioned considerations highlighted in this post.
Historically, scientists have shied away from using next-gen sequencing technology due to worries about successful library preparation and meaningful analysis. To address these concerns, Cofactor’s Project Scientists work with you to develop an experimental design that maximizes the data you obtain at a price you’re comfortable with. Our rigorous QC checkpoints and our extensive experience with library design and preparation (including low-input/limited material) help us achieve one of the highest success rates in the industry. And our ActiveSite interface helps you intuitively access and explore the significant amount of data generated by RNA-seq.
Overall, RNA-seq is a high-throughput, quantitive means of exploring the transcriptome of your organism of interest. Cofactor is excited and prepared to work with researchers in academic, government, or industrial labs to leverage our years of experience to design an experiment that will meet your needs. If you’re ready to get started on an RNA-seq project, we’re happy to help you reach your research goals. Are you and your team new to RNA-seq? Schedule a time for us to pay you a visit and provide an introduction to the technology and our approach. All you have to do is get in touch.
You might also be interested in how Mass Spectrometry stacks up to RNA-seq. Dr. Andy Benesh covers that in a later blog post.
To learn about profiling the immune system utilizing a new, multidimensional approach, this recorded talk might be helpful. The ImmunoPrism™ assay aligns with the drug development pipeline, from translational through clinical applications. Quantitative assessment of immune composition is important in understanding mechanism of action, confirming or predicting response, and developing predictive biomarkers to be used in a prognostic setting. ImmunoPrism enables you to leverage our investment in Health Expression Models, machine learning, and a highly-sensitive, validated molecular protocol. The assay is a complete solution – beginning with RNA extraction through immune profiling and onto the generation of multidimensional biomarkers. Each stage of the assay delivers an easy-to-interpret report, and our team would be happy to walk you through how you can make the most of the information you’ll receive.
Are you a fellow proponent of multidimensional data and biomarkers? We’d love to hear your take on what we can learn from others who are making strides in the field of Predictive Immune Modeling. Reach out to us anytime, using this form.
References:
https://eh.uc.edu/genomics/files/Illumina_Whitepaper_RNASeq_to_arrays_comparison.pdf


