In staying up to date with the latest trends in industry, the folks at Cofactor, myself included, often take a look at some popular blogs and other sources of information. Recently while browsing through RNA-seq Blog I came across an interesting poll they had put up concerning spike-in controls for RNA-seq experiments.
Now granted, a sample size of 16 voters (as of the time this poll was looked at) isn’t exactly a huge number from which to derive any conclusions, but it did get me thinking about the various opinions out there. Looking at this poll, it seems as if the majority of RNA-seq Blog’s readers-at least the ones that answered this poll question at this point in time-think that spike-in controls are important. Cofactor certainly believes spike-ins are important and we have implemented their use for at least a couple of different purposes. Which brings me to my next question: For those who believe in using spike-ins, why are they important to you? Are you using them for the robust quality control they provide? Or are you more interested in their use for confidently and correctly comparing expression levels of transcripts between samples? Perhaps both?
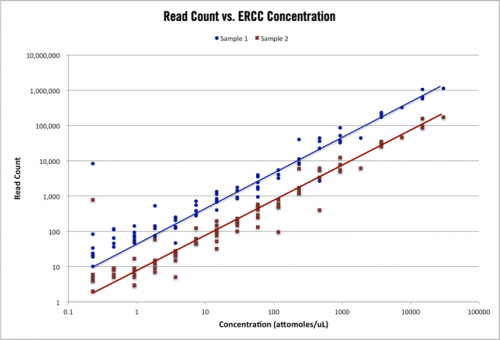
Cofactor is extremely excited about the work we’ve done in optimizing our spike-in controls and the confident results we’re providing to researchers through their use as part of our RNA-seq solution. In our eyes, the incorporation of spike-in controls is one of the key next steps in the evolution of RNA-seq, just as RNA-seq has by now largely replaced microarrays for comparative expression studies. Personally, I am very interested in seeing how the opinion on spike-in controls changes over the coming months as researchers become more familiar with the application and importance of this crucial molecular addition.